Journal of
eISSN: 2373-633X


Research Article Volume 14 Issue 1
1Department of Cellular and Molecular Sciences, Faculty of Advanced Sciences and Technology, Tehran Medical Sciences, Islamic Azad University, Iran
2Men’s Health and Reproductive Health Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran
3Skull Base Research Center, Loghman Hospital, Shahid Beheshti University of Medical Sciences, Iran
Correspondence: Farkhondeh Pouresmaeili, Men’s Health and Reproductive Health Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran, Tel +98- 23872572
Received: September 15, 2022 | Published: February 10, 2023
Citation: Eshghifar N, Rouhollah F, Barikrow N, et al. The expression analysis of long non-coding RNAs and p53 as a possible genetic signature in luminal: An invasive breast ductal carcinoma. J Cancer Prev Curr Res. 2023;14(1):9-13. DOI: 10.15406/jcpcr.2023.14.00511
Breast cancer is the second most common cause of cancer-related death among females. The long non-coding RNAs (lncRNAs) are a significant population of non-coding RNAs with well-defined functions in both adjacent normal cells and tumorigenesis. Miss expression of them has been associated with the development of different kinds of cancers. The aim of this study was to investigate the alterations in 3 lncRNAs and p53 in tissue samples of a group of women with luminal type A breast adenocarcinoma. In this case-control study, the expression levels of p53 and three lncRNAs were evaluated in association with luminal A breast cancer in 80 ductal carcinoma tumors and adjacent normal breast tissues. Quantitative real-time PCR was used to measure the expression of the mentioned genes. The data were analyzed using t-tests.
The expression levels of IGFBP7-AS1, and RHPN1-AS1, showed a significant increase (P> 0.05) while P53 and LINC00861 had a significant decrease in expression level in tumor tissues compared to adjacent normal tissues (P> 0.05). Receiver Operating Characteristic (ROC) curve indicated the diagnostic power of LINC00861 in differentiating tumor tissues from adjacent normal tissue with 90% sensitivity and 96% specificity, which can also interact with P53.
P53 and the LncRNAs including Linc00861, RHPN1-AS1, and IGFBP7-AS1 were dysregulated in invasive breast ductal carcinoma samples. Based on the area under the curve (AUC) value, expression of Linc00861, p53, RHPN1-AS1, and IGFBP-AS1 genes can help to differentiate patients from healthy individuals with diagnostic power of 0.855, 0.824, 0.778, and 0.659, respectively. The key roles of the latest genes in the control of cancer-related pathways and the dysregulation of their expression in studied malignancies imply that they can be exploited as biomarkers for cancer diagnosis, prognosis, and possible therapeutic targets. Here, the roles of lncRNAs in invasive breast ductal carcinoma type luminal A and their importance in prognosis and patient treatment are discussed.
Keywords: breast cancer, long non-coding RNA, prognostic biomarker
Breast cancer (BC) is one of the most frequent malignancies in women, globally.1 Although target therapies and surgery have improved the prognosis of BC patients in recent years, recurrence, metastasis, and drug resistance remain clinical problems that contribute to many BC patients' poor overall survival (OS).2-5 Thus, to enable breast cancer management and the development of patient outcomes, accurate and robust prognostic biomarkers and innovative therapeutic targets should be identified. LncRNAs have been indicated to be involved in mammary gland progression, as well as breast cancer development.6 They have been recommended as molecular biomarkers for the recognition of breast ductal carcinoma.7
According to their expression patterns and roles in Tumoral tissues, LncRNAs can be categorized as oncogenes and tumor suppressor genes (TSG).8 It has been demonstrated that they influence the plasticity of cancer stem cells. Based on the recent studies, the expression of RHPN1-AS1 is usually up-regulated in aggressive tumors, such as melanoma and non-small cell lung cancer.9,10 Furthermore, up-regulation of RHPN1-AS1 was notably associated with poor prognosis and malignancies; conversely, low expression of RHPN1-AS1 prevents metastasis and cancer cell proliferation.11
Several studies have found that, among many other cancers, IGFBP7 is implicated with a number of malignancies such as prostate cancer, hepatocellular carcinoma (HCC), gastroesophageal, breast, and colorectal cancers.12
P53 is the most important tumor suppressor genes. Mutations in this gene are most common in human cancers and induce tumor growth, increase angiogenesis, impair apoptosis, and resist therapy. Genetic alterations in P53 are associated with tumorigenesis, especially solid tumors such as breast, colon, and lung cancers.13,14
On the other hand, LINC00861 was implicated in the development and prognosis of a variety of cancer types. Low expression of LINC00861 was shown to be associated with advanced-stage cancers, poor survival, and lymph node metastases.15
Therefore, deciphering the function of lncRNAs in invasive breast ductal carcinoma could improve our knowledge of the molecular mechanisms underlying breast cancer progression.
The objective of this case-control study was to present a comprehensive overview of the relationship between the expression of P53, lncRNAs, and invasive breast ductal carcinoma type luminal A in Iranian population.
The Islamic Azad University, Tehran Medical Branch's ethical committee reviewed and approved all aspects of this project. Patients completed informed consent forms to allow the utilization of their specimens for research purposes. This study did not involve any additional costs for patients.
Patients and tissue specimens
80 fresh samples, 40 from luminal A tumors and 40 from adjacent normal tissues, were collected in liquid nitrogen from patients who were referred to Hamadan Hospital and maintained at -80°C until use for RNA extraction. The referring hospital pathologists characterized and confirmed the adjacent normal tissue specimens. None of the patients had received preoperative radiotherapy or chemotherapy. They also provided a signed informed consent form for study analysis. All protocols have been confirmed by Shahid Beheshti University of Medical Sciences (SBMU) Research Council ethic committee, and procedures have been carried out according to the appropriate instructions and regulations.
RNA extraction and RT-qPCR analysis
Total RNA was extracted from the fresh tissues using Trizol reagent (Thermo Fisher Scientific, Inc, USA) and quantified using Nanodrop 2000 according to the manufacturer's instructions for purity. The integrity of extracted nucleic acids was determined using agarose gel electrophoresis. Then, utilizing Random hexamers and Multiscribe reverse transcriptase (Applied Biosystems), based on manufacturer protocol, 1 ml of solution was added to the homogenized tissue sample. Then 200 μl of chloroform was added and incubated for 15-20 minutes at room temperature. Centrifugation was performed at 12000g at 4°C for 20 minutes. The supernatant was transferred to another microtube, 100% isopropanol was added with the same volume, after 10 minutes of incubation at room temperature and 15 minutes of centrifugation at 12000 g, the supernatant was discarded and 1 ml of 60% ethanol was added to the precipitate, and centrifuged at 7500 rpm. Then, the precipitate was dissolved in RNA-free water and stored at -80°C until usage. Agarose gel was used for the extracted RNA quality measurement and the adsorption rate was measured at 260 to 280 nm RNA using a nanodrop. In order to remove possible DNA contamination, the RNA reaction was treated by DNaseI before RT-PCR, the treated RNA was incubated with 0.5 μl of random hexamer enzyme for 5 min at 70°C. In the next step, 1 μl of dNTP mixture and 2 μl of RT buffer were added to the required amount of sterile distilled water, after 5 minutes of incubation at 37°C, 0.5 μl of RT enzyme was added. Incubation of the reaction mixture was performed at 42°C for 80 minutes.
Complementary DNA (cDNA) synthesis
For cDNA synthesis, about 500 ng of RNA with reverse transcriptase enzyme, oligo dt primers and random hexamer primers were placed in a final volume of 10 μl for 30 minutes at 37°C. It was placed at 80°C for 5 seconds in order to inactivate the enzyme. All steps were performed on ice and inside an aluminum rack.
The products were stored at - 20°C. Quantitative RT-PCR (Applied Biosystem 7500 Real‑Time PCR system- Thermo Fisher Scientific, Inc.) was performed to assess LncRNA expression levels utilizing SYBR Green PCR Master Mix and the StepOne Plus system with the following melting temperatures: 95 °C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 45 s.
In each experiment, a negative control was considered. Each sample was analyzed in duplicate. For standardization, as an internal control, the housekeeping gene, HPRT1 was selected; using the PCR designed primers by BLAST analysis of Genebank sequences which is listed as follows:
Gene name |
Primer sequences |
Linc00861 |
F: GGGAAATCAGAATACACAGT |
R: AGAGAGACAAGGAGCATC |
|
TP53 |
F: CTGTCATCTTCTGTCCCTTC |
R: TGGAATCAACCCACAGCTGCA |
|
IGFBP7-AS1 |
F:AGCAGCAGAAGACTATGA |
R:GTATGAGAGCAGGTGGTA |
|
RHPN1-AS1 |
F: GCTCCTGGTCATCAAGTTCCTCT |
R: GCACAGGCACCAGAATGATCC |
|
HPRT1 |
F:AGCCTAAGATGAGAGTTC |
|
R:CACAGAACTAGAACATT |
Statistical analysis
Statistical analysis for all experiments was performed using the SPSS version 24.0 (SPSS, Chicago, IL, USA). Statistical data are presented as the Bayesian estimation supersedes the t-test to determine the significant differences in means between the expression levels of cancerous cells and adjacent normal tissues. The survival analysis was performed utilizing a Kaplan‑Meier test to evaluate the statistical significance between the two groups. P<0.05 was considered to demonstrate a statistically significant difference. For parameters, a t student prior family with 4000 iterations and 2000 burn-outs was considered. The P values were calculated from the Frequentist technique including the median test. The Spearman method was performed to evaluate the correlation between relative gene expressions. To implement analysis in the software R 4.03R, jags, and ggplot2 packages were used. P-value <0.05 was considered statistically significant.
In this study, the demographic information of the patients, as well as their clinicopathological information was considered (Table 1). RNA was extracted from the samples and was quantified at 260/280 with ratios between 1.7 and 1.9. The relative expression levels of the stated genes as possible diagnostic biomarkers were evaluated by Real Time Quantitative PCR. Receiver Operating Characteristic (ROC) curve including all criteria were created. The area under the curve (AUC) which is a notable number to demonstrate the diagnostic accuracy of the gene expression level was measured.
Parameters |
Values |
Age [mean±SD, (range)] |
51.82±11.25 (25-32) |
Site of primary tumor |
|
Right breast |
21 (52.5%) |
Left breast |
19 (47.5%) |
Cancer stage (%) |
|
I |
2 (5%) |
II |
24 (60%) |
III |
14 (35%) |
Overall grade (%) |
|
I |
5 (12.5%) |
II |
19 (47.5%) |
III |
14 (35%) |
Unknown |
2 (5%) |
Lymphatic invasion |
|
Yes |
32 (80%) |
No |
8(20%) |
Vascular invasion |
|
Yes |
33 (82.5%) |
No |
7 (17.5%) |
Tumor size (%) |
|
≤2 cm |
5 (12.5%) |
>2 |
35 (87.5%) |
Estrogen receptor (%) |
|
Positive |
13 (32.5%) |
Negative |
4 (10%) |
Unknown |
23 (57.5%) |
Progesterone receptor (%) |
|
Positive |
11 (27.5%) |
Negative |
4 (10%) |
Unknown |
25 (62.5%) |
Her2/neu expression (%) |
|
Positive |
4 (10%) |
Negative |
14 (35%) |
Unknown |
22 (55%) |
Table 1 Patients’ demographic information
This result ranges from 0 to 1, with 1 defined as a flawlessly exact test and 0.5 concerning no discrimination. The values greater than 0.9, were regarded as outstanding.
Co-expression of any of examined LncRNAs and P53 was considerably associated with metastasis (Table 2). According to t-Test analysis, the expression of all LncRNAs was significantly different in two groups of tissue samples (Table 3).
Parameters |
P53 up-regulation |
P53 down-regulation |
P value |
Age |
0.72 |
||
<55 |
11 (55%) |
9 (45%) |
|
≥55 |
9 (45%) |
11 (55%) |
|
Site of primary tumor |
0.64 |
||
Right breast |
10 (47%) |
11 (53%) |
|
Left breast |
10 (52%) |
9 (48%) |
|
Stage |
0.03 |
||
1 |
0 (0%) |
2 (100%) |
|
2 |
12 (50%) |
12 (50%) |
|
3 |
7 (50%) |
7 (50%) |
|
Histological Grade |
0. 54 |
||
1 |
3 (60%) |
2 (40%) |
|
2 |
11 (57%) |
8(43%) |
|
3 |
6(43%) |
8 (57%) |
|
Lymphatic invasion |
0.69 |
||
Yes |
16 (50%) |
16 (50%) |
|
No |
7 (87.5%) |
1 (12.5%) |
|
Vascular invasion |
0.48 |
||
Yes |
15 (45%) |
18 (55%) |
|
No |
6 (85%) |
1 (15%) |
|
Tumor size |
0.8 |
||
≤2 |
4 (80%) |
1 (20%) |
|
>2 |
18 (51%) |
17 (49%) |
|
ER status |
0.69 |
||
Positive |
4 (31%) |
9 (69%) |
|
Negative |
4 (100%) |
0 (0%) |
|
PR status |
0.1 |
||
Positive |
6 (54 %) |
5(46%) |
|
Negative |
3 (75%) |
1 (25%) |
|
Her2 status |
0.9 |
||
Positive |
2 (50%) |
2 (50%) |
|
Negative |
8 (57%) |
6 (43%) |
|
Table 2 The expression level of P53 is shown in relation to the participants’ parameters
Gene |
Posterior mean diff. |
SD |
Effect size |
P-value |
95% HDI |
IGFBP7-AS1 |
-1.652 |
3.41 |
-0.497 |
0.004 |
[-2.99, -0.38] |
LINC00861 |
-4.886 |
3.45 |
-1.499 |
<0.0001 |
[-6.18, -3.51] |
P53 |
-3.184 |
3.6 |
-0.914 |
<0.0001 |
[-4.46, -1.87] |
RHPN-AS1 |
-1.861 |
2.06 |
-0.932 |
<0.0001 |
[-2.66, -1.08] |
Table 3 The comparison relative gene expression between tumor and healthy samples; Results of Bayesian estimation supersedes the t-test
Abbreviations: SD, standard deviation; HDI, highest density interval
LINC00861 is implicated in a significant biological mechanism of invasive breast ductal carcinoma; a possible correlation between LINC00861 and clinical biomarkers was discovered. Its level in tumor tissue was significantly down-regulated compared to adjacent normal tissue. The results of this study indicated that Linc00861 could play a role as a tumor suppressant in the progression of invasive luminal type A breast cancer. Furthermore, there was no correlation between LINC00861 levels and differentiation, tumor size, age, and, histology appearance (Table 3). According to the data, patients with low expression of LINC00861 had shorter survival than patients with high expression ones (Figure 1).
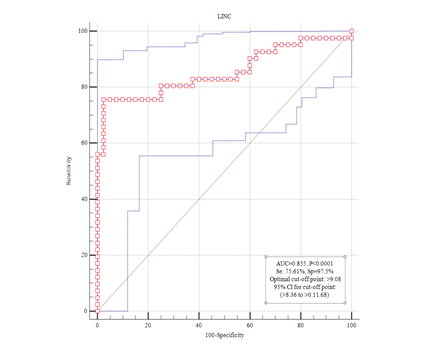
Figure 1 Kaplan – Meier curve (KM). Correlation between linc00861 expression and prognosis of invasive ductal carcinoma luminal type A. This curve indicates the accuracy of linc00861 in distinguishing between turmeric and healthy tissue.
RHPN1-AS1 expression was up regulated in BC which was substantially associated with a poor prognosis of breast cancer (Figure 2). Furthermore, RHPN1-AS1 was found to be an independent and remarkable possible predictor of breast cancer prognosis in both univariate and multivariate studies. These data imply that RHPN1-AS1 might be a predictive biomarker and therapeutic target for breast cancer.
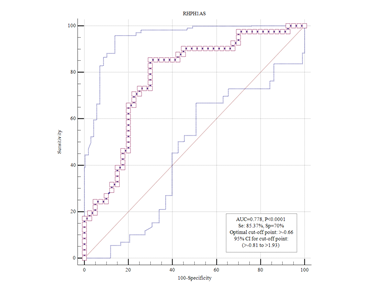
Figure 2 Kaplan – Meier curve (KM). Correlation between RHPN1-AS1 expression and prognosis of invasive ductal carcinoma luminal type A. This curve indicates the accuracy of RHPN1-AS1 in distinguishing between turmeric and healthy tissue.
In the present study, it was shown that IGFBP7-AS1 was up-regulated in tumor tissues in comparison with the adjacent normal cells. This change in expression was not significantly associated with lymph node metastasis and age (Figure 3). Since IGFBP7-AS1 has mitogenic and anti-apoptotic effects, it might be increased the risk of breast cancer. Therefore, this biomarker can be used as a prognostic indicator in luminal type A invasive breast ductal carcinoma.
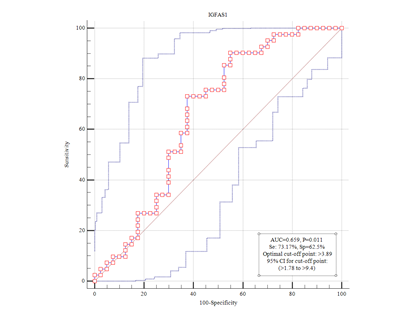
Figure 3 Kaplan – Meier curve (KM). Correlation between IGFBP7-AS1 expression and prognosis of invasive ductal carcinoma luminal type A. This curve indicates the accuracy of IGFBP7-AS1 in distinguishing between turmeric and healthy tissue.
Also, In comparison with adjacent normal tissue, the low expression level of p53 in tumor samples was detected (Figure 4). This change in p53 gene expression was associated with a poor prognosis in invasive breast ductal carcinoma. No significant correlation between age, tumor size, and p53 expression level was observed.
The abnormal expression of lncRNAs has been significantly implicated in a variety of cancers, including colorectal16 cervical cancer,17 and prostate.18 Numerous researches provided evidence that lncRNAs influence cancer, such as invasive breast cancer, acting as either tumor suppressors or oncogenes.19 Despite the great development in Breast cancer therapeutic techniques recently, the prognosis of invasive breast cancer patients remains unknown.20-22 In invasive breast ductal carcinoma, low expression of LINC00861 was associated with a poor prognosis. Also, evidence has indicated that LINC00861 is involved in the progression and occurrence of different kinds of cancers. The differential expression of LINC00861 among non-early and early recurrence in patients with hepatocellular carcinoma might be utilized as a biomarker to predict the early recurrence of liver cancer following curable resection.23
It has been reported that the low expression of LINC 00861 is associated with a poor prognosis in ovarian cancer.24 Linc00861 can be an important biomarker for the early diagnosis and treatment of the disease. The results of the present study indicated that LINC00861 can interact with P53. LINC00861 may control basal biological processes and may be used as a potential clinical biomarker for diagnosis. Since the expression of lncRNAs could serve as oncogene and tumor suppressor, the miss-regulation of their expression is associated with a poor prognosis in patients.
RHPN1-AS1 is involved in cell proliferation, migration, and promotion of cell invasion. In the study of Lu et al., An increase in RHPN1-AS1 expression was reported in the melanoma cancer cell line.10 In contrast, another study in lung cancer found that the expression of RHPN1-AS1 in tumor tissue was down regulated in comparison with healthy tissue, which indicates the suppressive role of RHPN1-AS1.9 Xu et al., reported that low expression of RHPN1-AS1 suppresses the proliferation and metastasis of cancer cells.11 These observations suggest that, depending on the type of cancer, RHPN1-AS1 may play an oncogenic or suppressive role.
On the other hand, P53 is one of the most significant therapeutic targets for cancer treatment due to its critical functions in carcinogenesis prevention. Various treatments can be performed with combination of tumor and adjacent normal cells, based on the type of genetic variation in P53.
It seems that measuring the level of IGFBP7-AS1 in breast tissue can be used as a prognostic indicator for the diagnosis of luminal type A breast cancer. Also, due to the tumor genesis of the IGF system, drug interactions for this system are being studied and various strategies in this field can be effective in the treatment of this type of cancer.
In addition, by examining the signaling pathways involved more comprehensively, new upstream and downstream proteins that function as targets can be discovered.
The identification of stage, subtype or even metastasis lncRNA expression in invasive breast ductal carcinoma may make them fruitful biomarkers for efficient prognosis and diagnosis. The progress of RNA-based treatment methods has opened up these lncRNAs as new targets for targeting breast cancer for those breast cancer-related lncRNAs whose function has been experimentally verified.
F.P.; project design, co-wrote the paper and critical revision of the manuscript for important intellectual content; M.T; Study concept and design, Acquisition of data; collecting samples, N.E; Acquisition of data, performed the searches ,co-wrote the paper, Performed proof editing,; F.R: Analysis and interpretation of data, Critical revision of the manuscript for important intellectual content. N.B: Critical revision of the manuscript for important intellectual content, Performed proof editing. All the authors provided their final approval for the completed manuscript.
None.
The authors declare no conflicts of interest.

©2023 Eshghifar, et al. This is an open access article distributed under the terms of the, which permits unrestricted use, distribution, and build upon your work non-commercially.